by Tapiwa Tarwa
For decades, the medical field has waged war against disease-causing bacteria. Armed with the (then) recent discovery of antimicrobials, medical professionals finally believed they had the upper hand. However; these bacteriae have proven to be a worthy opponent as they have quickly retaliated with antimicrobial resistance (AMR)*. This resistance is a weapon rendering nearly all antimicrobials ineffective. This leaves the health system defenceless against the persistence and spread of many deadly diseases. As the medical field continues to lose ground in the war against disease, it becomes more critical to understand the vast arsenal of tools at the bacteriae’s disposal – in particular, those that allow them to become resistant.
- Antimicrobial resistance- the ability to resist the effects of antimicrobials that could successfully treat the bacteria
- Sequencing- a process of determining the DNA (genetics) of an organism
- Niche- a comfort place for life
- A battery of tests- a series of tests
Recent advances in the medical science field, especially those in sequencing*, have shed light on the biological properties that are altered (for example, through mutations) to allow for resistance. However, these advances fail to explain how such resistance occurs or spreads within a population during treatment. Researchers from Princeton University set out to mimic bacterial niches* and show the precise molecular mechanism that underpins the rapid emergence of antimicrobial resistance in bacteria [1,2].
In a two-paper series, the researchers conducted a battery* of experiments using many techniques to further the understanding of resistance. The main techniques included a microfluidic system (a technology that allows for precise manipulation of physiochemical conditions in bacterial cultures- temperature, nutrients, antibiotic concentrations, etc.) and time-lapse microscopy (series of images viewed at a higher speed to give an accelerated view of the microscopic process) [1,2].
In the first paper, the researchers used the microfluidic system to expose several E. coli to ciprofloxacin (a genotoxic antimicrobial) concentration gradient. This concentration gradient mimicked those seen in patients on treatment as it is often difficult to obtain adequate amounts of antibiotics at infection sites due to their location. This experiment turned the current understanding of resistance on its head [1].
Resistant bacteria were thought to be already present in large populations of bacteria (i.e. 1 in 108). However, this experiment showed that resistant bacteria were not pre-existing in a sizeable bacterial population (a subpopulation) but were instead acquired as de novo mutations due to antimicrobial exposure (Figure 1) [1].
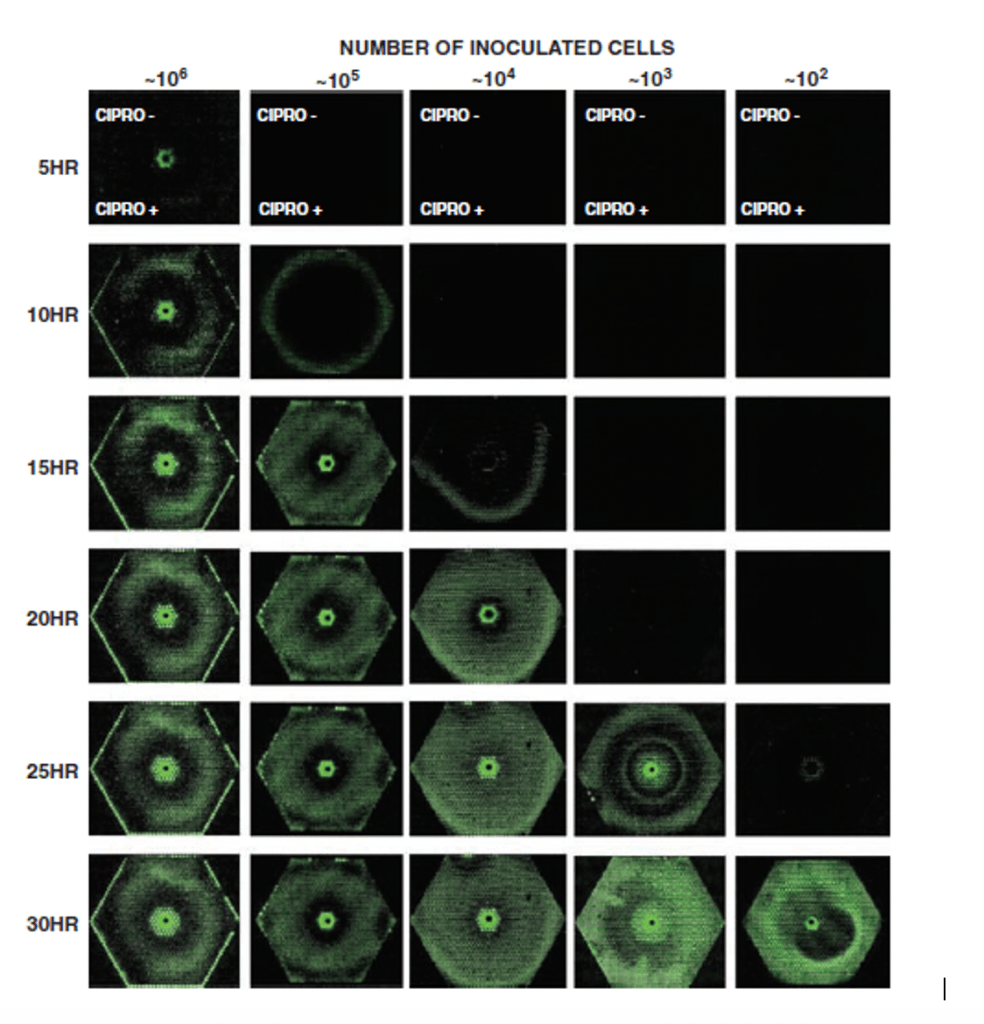
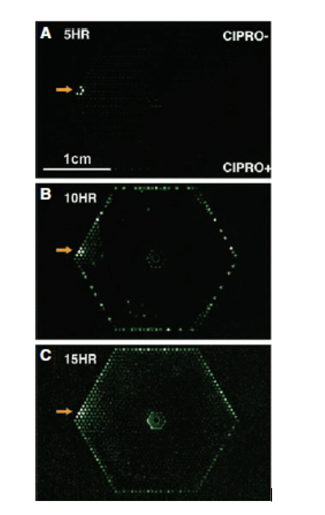
MIC- Minimum inhibitory concentration -the lowest concentration of an antimicrobial that will inhibit bacterial growth
Additionally, the filamentous bacteria seemed to produce multiple chromosomes (polyploidy). The induction of the SOS response (a response traditionally thought to be a DNA repair mechanism) is required for the formation of filaments and multiple chromosomes. However, the researchers suggest that rather than stopping the cell cycle for repair, the SOS response engages a low fidelity polymerase[1] that continues to copy the chromosome generating mutations. The mutant chromosomes are then supposedly passed to the resistant progeny (Figure 4) [2].
Polymerase- an enzyme that is essential in replicating DNA

What does this all mean in the fight against bacteria? Well, it seems as though resistance is in their genes and they will not go down without a fight. As the medical field scramble for more weapons and tools to fight this worthy opponent, one thing is for sure- “what doesn’t kill the bacteria only makes them stronger.” Thus, future interventions should not only aim to win the battle but rather the war.

- Zhang Q, Lambert G, Liao D, Kim H, Robin K, Tung CK, Pourmand N, Austin RH. Acceleration of emergence of bacterial antibiotic resistance in connected microenvironments. Science. 2011 Sep 23;333(6050):1764-7.
- Bos J, Zhang Q, Vyawahare S, Rogers E, Rosenberg SM, Austin RH. The emergence of antibiotic resistance from multinucleated bacterial filaments. Proceedings of the National Academy of Sciences. 2015 Jan 6;112(1):178-83.
- Baker S, Duy PT, Nga TV, Dung TT, Phat VV, Chau TT, Turner AK, Farrar J, Boni MF. Fitness benefits in fluoroquinolone-resistant Salmonella Typhi in the absence of antimicrobial pressure. Elife. 2013 Dec 10;2: e01229.
Leave a comment